class: title-slide, inverse, center, middle # Generalized Additive Models <div style="position: absolute; bottom: 15px; vertical-align: center; left: 10px"> <img src="images/02-foundation-vertical-white.png" height="200"> </div> --- # Overview - What is a GAM? - What is smoothing? - How do GAMs work? - Fitting GAMs using `dsm` --- class: inverse, middle, center # What is a GAM? --- # "gam" 1. *Collective noun used to refer to a group of whales, or rarely also of porpoises; a pod.* 2. *(by extension) A social gathering of whalers (whaling ships).* (via Natalie Kelly, AAD. Seen in Moby Dick.) --- # Generalized Additive Models - Generalized: many response distributions - Additive: terms **add** together - Models: well, it's a model... --- # What does a model look like? - Count `\(n_j\)` distributed according to some count distribution - Model as sum of terms 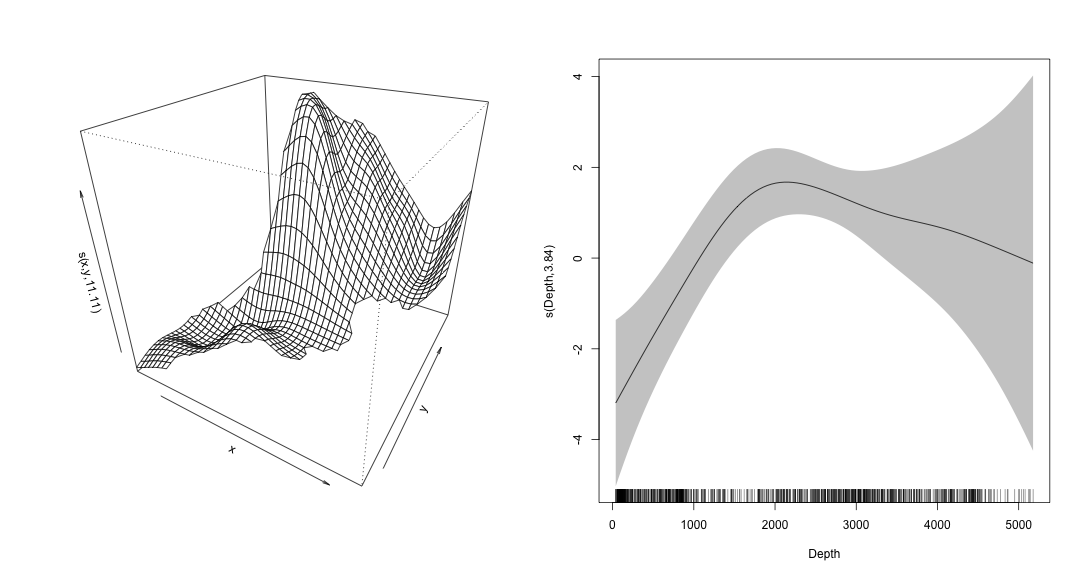<!-- --> --- # Mathematically... Taking the previous example... $$ n_j = \color{red}{A_j}\color{blue}{\hat{p}_j} \color{green}{\exp}\left[\color{grey}{ \beta_0 + s(\text{y}_j) + s(\text{Depth}_j)} \right] + \epsilon_j $$ where `\(\epsilon_j\)` are some errors, `\(\quad n_j\sim\)` count distribution - `\(\color{red}{\text{area of segment - offset}}\)` - `\(\color{blue}{\text{probability of detection in segment}}\)` - `\(\color{green}{\text{link function}}\)` - `\(\color{grey}{\text{model terms}}\)` --- # Response $$ \color{red}{n_j} = A_j\hat{p}_j \exp\left[ \beta_0 + s(\text{y}_j) + s(\text{Depth}_j) \right] + \epsilon_j $$ <br/> where `\(\epsilon_j\)` are some errors, `\(\quad \color{red}{n_j\sim \text{count distribution}}\)` --- # Count distributions .pull-left[ 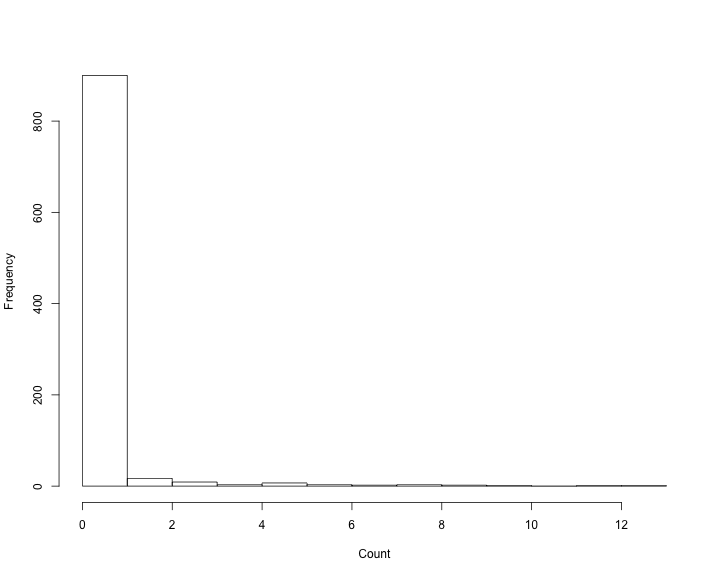<!-- --> ] .pull-right[ - Response is a count (not not always integer) - Often, it's mostly zero (that's complicated) - Want response distribution that deals with that - Flexible mean-variance relationship ] --- # Tweedie distribution .pull-left[ 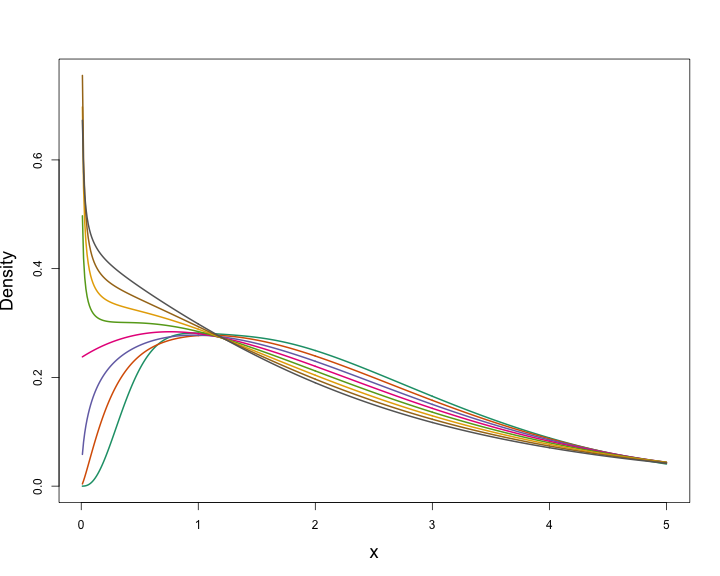<!-- --> ] .pull-right[ - `\(\text{Var}\left(\text{count}\right) = \phi\mathbb{E}(\text{count})^q\)` - Common distributions are sub-cases: - `\(q=1 \Rightarrow\)` Poisson - `\(q=2 \Rightarrow\)` Gamma - `\(q=3 \Rightarrow\)` Normal - We are interested in `\(1 < q < 2\)` - (here `\(q = 1.2, 1.3, \ldots, 1.9\)`) ] --- # Negative binomial distribution .pull-left[ 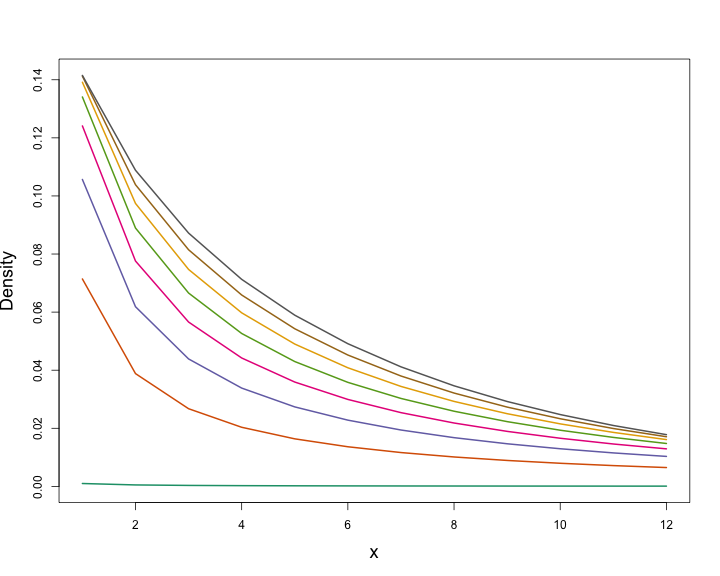<!-- --> ] .pull-right[ - `\(\text{Var}\left(\text{count}\right) =\)` `\(\mathbb{E}(\text{count}) + \kappa \mathbb{E}(\text{count})^2\)` - Estimate `\(\kappa\)` - Is quadratic relationship a "strong" assumption? - Similar to Poisson: `\(\text{Var}\left(\text{count}\right) =\mathbb{E}(\text{count})\)` ] --- # Smooth terms $$ n_j = A_j\hat{p}_j \exp\left[ \beta_0 + \color{red}{s(\text{y}_j) + s(\text{Depth}_j}) \right] + \epsilon_j $$ <br/> where `\(\epsilon_j\)` are some errors, `\(\quad n_j\sim\)` count distribution --- # Okay, but what about these "s" things? .pull-left[ <!-- --> ] .pull-right[ - Think `\(s\)`=**smooth** - Want to model the covariates flexibly - Covariates and response not necessarily linearly related! - Want some wiggles ] --- class: inverse, middle, center # What is smoothing? --- # Straight lines vs. interpolation .pull-left[ <!-- --> ] .pull-right[ - Want a line that is "close" to all the data - Don't want interpolation -- we know there is "error" - Balance between interpolation and "fit" ] --- # Splines .pull-left[ <!-- --> ] .pull-right[ - Functions made of other, simpler functions - **Basis functions** `\(b_k\)`, estimate `\(\beta_k\)` - `\(s(x) = \sum_{k=1}^K \beta_k b_k(x)\)` - Makes the maths much easier ] --- # Measuring wigglyness - Visually: - Lots of wiggles == NOT SMOOTH - Straight line == VERY SMOOTH - How do we do this mathematically? - Derivatives! - (Calculus *was* a useful class after all) --- # Wigglyness by derivatives  --- # Making wigglyness matter - Integration of derivative (squared) gives wigglyness - Fit needs to be **penalised** - **Penalty matrix** gives the wigglyness - Estimate the `\(\beta_k\)` terms but penalise objective - "closeness to data" + penalty --- # Penalty matrix - For each `\(b_k\)` calculate the penalty - Penalty is a function of `\(\beta\)` - `\(\lambda \beta^\text{T}S\beta\)` - `\(S\)` (*penalty matrix*) calculated once - *smoothing parameter*, `\(\lambda\)` dictates influence --- # Smoothing parameter <!-- --> --- # How wiggly are things? - We can set **basis complexity** or "size" `\(k\)` - Maximum wigglyness - Smooths have **effective degrees of freedom** (EDF) - EDF < `\(k\)` - Set `\(k\)` "large enough" --- # Why GAMs are cool... .pull-left[  ] .pull-right[ - Fancy smooths (cyclic, boundaries, ...) - Fancy responses (exp family and beyond!) - Random effects (by equivalence) - Markov random fields - Correlation structures - See Wood (2006/2017) for a handy intro ] --- class: inverse, middle, center # Okay, that was a lot of theory... --- class: inverse, middle, center # Example data --- # Example data <img src="images/data_ships.png"> --- # Example data <img src="images/observers.png"> --- # Sperm whales off the US east coast .pull-left[ <img src="images/spermwhale.png" width="100%"> ] .pull-right[ - Hang out near canyons, eat squid - Surveys in 2004, US east coast - Combination of data from 2 NOAA cruises - Thanks to Debi Palka (NOAA NEFSC), Lance Garrison (NOAA SEFSC) for data. Jason Roberts (Duke University) for data prep. ] --- # Model formulation - Pure spatial, pure environmental, mixed? - May have some prior knowledge - Biology/ecology - What are drivers of distribution? - Inferential aim - Abundance - Ecology --- class: inverse, middle, center # Fitting GAMs using dsm --- # Translating maths into R $$ n_j = A_j\hat{p}_j \exp\left[ \beta_0 + s(\text{y}_j) \right] + \epsilon_j $$ <br/> where `\(\epsilon_j\)` are some errors, `\(\quad n_j\sim\)` count distribution <br/> - inside the link: `formula=count ~ s(y)` - response distribution: `family=nb()` or `family=tw()` - detectability: `ddf.obj=df_hr` - offset, data: `segment.data=segs, observation.data=obs` --- # Your first DSM ```r library(dsm) dsm_x_tw <- dsm(count~s(x), ddf.obj=df, segment.data=segs, observation.data=obs, family=tw()) ``` `dsm` is based on `mgcv` by Simon Wood --- # What did that do? ```r summary(dsm_x_tw) ``` ``` ## ## Family: Tweedie(p=1.326) ## Link function: log ## ## Formula: ## count ~ s(x) + offset(off.set) ## ## Parametric coefficients: ## Estimate Std. Error t value Pr(>|t|) ## (Intercept) -19.8115 0.2277 -87.01 <2e-16 *** ## --- ## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 ## ## Approximate significance of smooth terms: ## edf Ref.df F p-value ## s(x) 4.962 6.047 6.403 1.07e-06 *** ## --- ## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 ## ## R-sq.(adj) = 0.0283 Deviance explained = 17.9% ## -REML = 409.94 Scale est. = 6.0413 n = 949 ``` --- # Plotting .pull-left[ <!-- --> ] .pull-right[ - `plot(dsm_x_tw)` - Dashed lines indicate +/- 2 standard errors - Rug plot - On the link scale - EDF on `\(y\)` axis ] --- # Adding a term - Just use `+` ```r dsm_xy_tw <- dsm(count ~ s(x) + s(y), ddf.obj=df, segment.data=segs, observation.data=obs, family=tw()) ``` --- # Summary ```r summary(dsm_xy_tw) ``` ``` ## ## Family: Tweedie(p=1.306) ## Link function: log ## ## Formula: ## count ~ s(x) + s(y) + offset(off.set) ## ## Parametric coefficients: ## Estimate Std. Error t value Pr(>|t|) ## (Intercept) -20.0908 0.2381 -84.39 <2e-16 *** ## --- ## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 ## ## Approximate significance of smooth terms: ## edf Ref.df F p-value ## s(x) 4.943 6.057 3.224 0.004239 ** ## s(y) 5.293 6.420 4.034 0.000322 *** ## --- ## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 ## ## R-sq.(adj) = 0.0678 Deviance explained = 27.4% ## -REML = 399.84 Scale est. = 5.3157 n = 949 ``` --- # Plotting ```r plot(dsm_xy_tw, pages=1) ``` <!-- --> - `scale=0`: each plot on different scale - `pages=1`: plot together --- # Bivariate terms - Assumed an additive structure - No interaction - We can specify `s(x,y)` (and `s(x,y,z,...)`) --- # Thin plate regression splines - Default basis - One basis function per data point - Reduce # basis functions (eigendecomposition) - Fitting on reduced problem - Multidimensional --- # Thin plate splines (2-D) <img src="images/tprs.png" alt="Thin plate regression spline basis functions. Taken from Wood 2006."> --- # Bivariate spatial term ```r dsm_xyb_tw <- dsm(count ~ s(x, y), ddf.obj=df, segment.data=segs, observation.data=obs, family=tw()) ``` --- # Summary ```r summary(dsm_xyb_tw) ``` ``` ## ## Family: Tweedie(p=1.29) ## Link function: log ## ## Formula: ## count ~ s(x, y) + offset(off.set) ## ## Parametric coefficients: ## Estimate Std. Error t value Pr(>|t|) ## (Intercept) -20.2745 0.2477 -81.85 <2e-16 *** ## --- ## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 ## ## Approximate significance of smooth terms: ## edf Ref.df F p-value ## s(x,y) 16.89 21.12 4.333 3.73e-10 *** ## --- ## Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1 ## ## R-sq.(adj) = 0.102 Deviance explained = 34.7% ## -REML = 394.86 Scale est. = 4.8248 n = 949 ``` --- # Plotting... erm... .pull-left[ <!-- --> ] .pull-right[ ```r plot(dsm_xyb_tw) ``` ] --- # Let's try something different .pull-left[ ```r plot(dsm_xyb_tw, select=1, scheme=2, asp=1) ``` - Still on link scale - `too.far` excludes points far from data ] .pull-right[ <!-- --> ] --- # Comparing bivariate and additive models <!-- --> --- class: inverse, middle, center # Let's have a go...